|
Users can perform simple and advanced searches based on annotations relating to sequence, structure and function. The RCSB PDB also provides a variety of tools and resources. Dictionary of Secondary Structure of Proteins (DSSP) assigns eight state secondary structure using hydrogen bonds alone. Janes, 2010, 2Struc - The Protein Secondary Structure Analysis Server, Biophysical Journal, 98:454a-455) and each of the methods you run. Free for commercial use High Quality Images. If you use 2Struc and publish your work please cite our paper (Klose, D & R.W. 71000+ Vectors, Stock Photos & PSD files. You can find data in Pfam in various ways. As a member of the wwPDB, the RCSB PDB curates and annotates PDB data according to agreed upon standards. Find & Download Free Graphic Resources for Protein Structure. all UniProt and NCBI GI) or different levels of redundancy. In addition to protein secondary structure, JPred also makes predictions. Pfam full alignments are available from searching a variety of databases, either to provide different accessions (e.g. Other sites for secondary structure predictions include: JPred4 - is the latest version of the popular JPred protein secondary structure prediction server which provides predictions by the JNet algorithm, one of the most accurate methods for secondary structure prediction. The data presented for each entry is based on the UniProt Reference Proteomes but information on individual UniProtKB sequences can still be found by entering the protein accession. Based primarily on hydrogen bonding patterns and some geometric constraints, it assigns every residue to one of eight possible states. 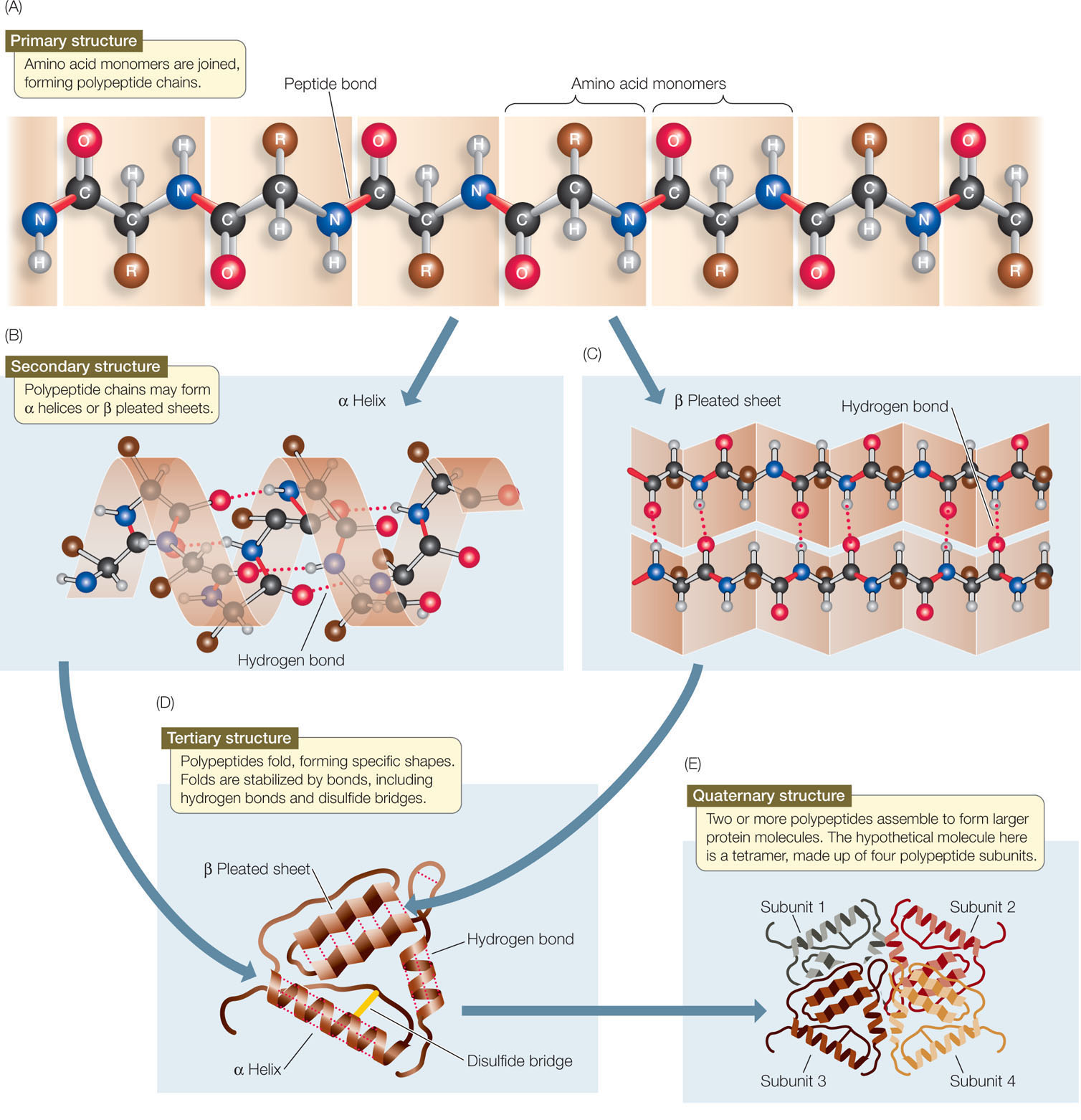
A clan is a collection of Pfam entries which are related by similarity of sequence, structure or profile-HMM. Define Secondary Structure of Proteins (DSSP) is the standard tool for the annotation of secondary structure elements from protein structures ( Kabsch and Sander, 1983 Touw et al., 2015 ). Pfam also generates higher-level groupings of related entries, known as clans. The identification of domains that occur within proteins can therefore provide insights into their function. Different combinations of domains give rise to the diverse range of proteins found in nature. Proteins are generally composed of one or more functional regions, commonly termed domains.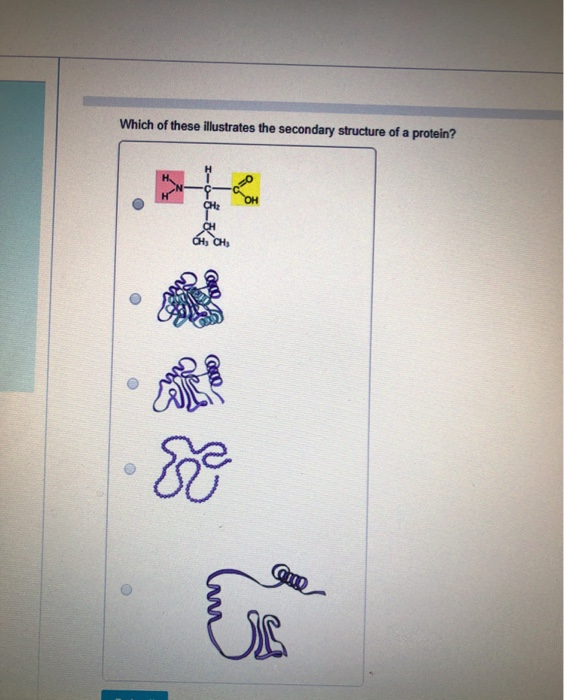 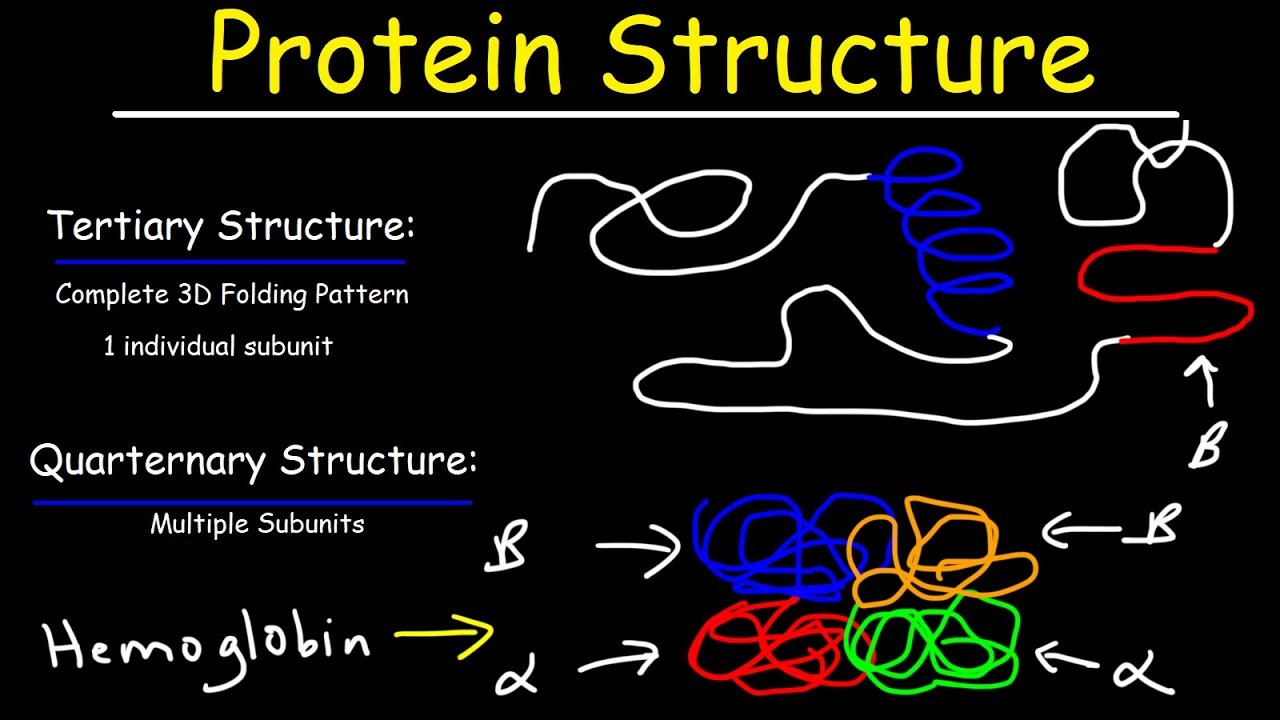  
The Pfam database is a large collection of protein families, each represented by multiple sequence alignments and hidden Markov models (HMMs).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
